Areas of Focus
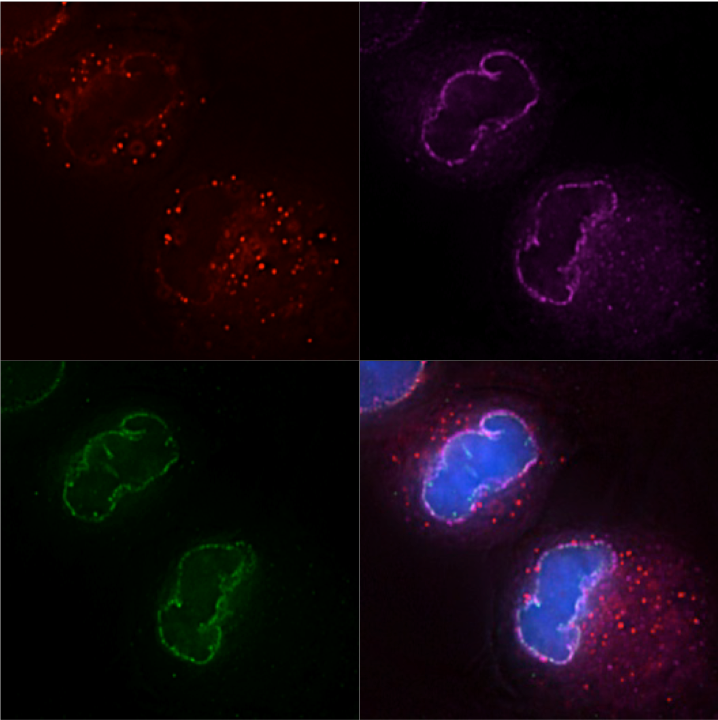
HIV-1, Mx2, and the nuclear pore complex
How does nuclear pore heterogeneity affect the antiviral activity of Mx2?
HIV-1 accesses the nuclear DNA of interphase cells via a poorly defined process involving functional interactions between the capsid protein (CA) and nucleoporins (Nups). Nups also mediate the function of the antiviral protein Mx2, which inhibits nuclear import of the HIV-1 pre-integration complex. We have observed striking variability in Nup-dependent HIV-1 nuclear import and Mx2 activity depending on the cell-type, cell-cycle, and CA-cyclophilin A (CypA) interactions. Our preliminary data indicate that compositional and/or conformational heterogeneity in nuclear pores both within and between cells can have dramatic effects on HIV-1 infection. This heterogeneity also alters the antiviral activity of Mx2. Previous investigations have relied on methods that dramatically alter cellular nuclear import, but new technologies now allow us to manipulate the functional capability of nuclear pores without disrupting pore formation. We are interested in using these precise manipulations to define the Nup-CA and Nup-Mx2 interactions that are functionally relevant to HIV-1 infection and in observing how the virus adapts to changes in available nuclear import pathways.
Genetic basis for retrovirus resistance
What are the genes responsible for protective antiviral responses in retrovirus resistant individuals?
The vast majority of individuals fail to mount an effective immune response against retroviral infection and eventually succumb to virally-induced disease. However, some individuals are able to mount neutralizing immune responses that are robust, long-lasting, and virus-neutralizing. Inbred mice are a powerful tool for the identification of genes that control antiviral immune responses since individuals of a given strain are genetically identical and genes can be definitively linked to resistant phenotypes, which is often not possible in human genetic studies. We utilize classical genetics in combination with genomic approaches to identify genes that determine resistance or susceptibility to retroviral infections in mice in order to better understand the natural genetic variability that may affect the outcome of viral infections in humans. We are currently investigating a recessive locus responsible for a novel pathway underlying the generation of antiviral antibodies in murine leukemia virus infected mice.
Antiretroviral immune responses
What are the signaling pathways underlying neutralizing antiretroviral immune responses in resistant individuals?
Virus-specific immune responses are initiated by the recognition of pathogen associated molecular patterns (PAMPs) by pattern recognition receptors (PRRs). This detection results in the activation of innate immune effectors, stimulation of inflammatory responses, and directs the pathogen-specific adaptive response. We utilize inbred mice to investigate the requirements for antiviral immunity in vivo since this allows for the investigation of the immune response to retroviral infections in their natural host. Furthermore, the canonical pathway for anti-retroviral antibody production in mice is TLR7 and interferon-𝝲 (IFN𝝲) dependent, however, we have recently identified an IFN𝝲-independent pathway for antiviral antibody production and class-switch recombination; and a TLR7-independent pathway for neutralizing antibody production. We are interested in identifying the cell types and signaling networks underlying these unique pathways using genetic, immunological, and molecular biology approaches.
Intrinsic cellular defenses
How do intrinsic innate immune effectors restrict retroviral replication?
An essential component of host defense against viral infection is the arsenal of proteins with direct antiviral activity that target specific steps in the viral replication cycle. These proteins function as an intrinsic immune system, or as a specialized component of the innate immune system, and are often termed “restriction factors”. Importantly, such proteins are able to dominantly and autonomously exhibit antiviral activity, including in cell culture assays. The antagonistic coevolution of virus and host selects for adaptations that alter the specificity and/or function of these antiviral proteins, and variation in intrinsic immunity is a determinant of viral tropism of similar importance to the variation in host cell factors required for viral replication. We are interested in understanding the mechanism by which these factors restrict the replication of retroviruses, and in the determinants for species-specificity. Currently, we are particularly interested in genes that specifically target the replication of foamy viruses.